Bottom
Given the role of spA as a fundamental virulence factor decisive for the ability of Staphylococcus aureus to escape innate and adaptive immune responses, it can be considered as an object subject to adaptive evolution and that variations in Staphylococcus aureus (SpA) Recombinants can uncover variations in pathogenicity.
Results
The genetic structure of the population was inferred from the extracellular domains of the SpA gene sequence (AE domains and the X region) and compared with MLST analysis of 41 genetically diverse methicillin-resistant (MRSA)-sensitive S. aureus. methicillin (MSSA). are. The inconsistency between tree topologies was notable and in the inferred spA tree, most of the MSSA isolates clustered in a distinct group. In contrast, the distribution of the strains according to their spA type was not always consistent with the inferred tree of the complete spA gene, predicting that spA is a mosaic gene composed of different segments that exhibit different evolutionary histories.
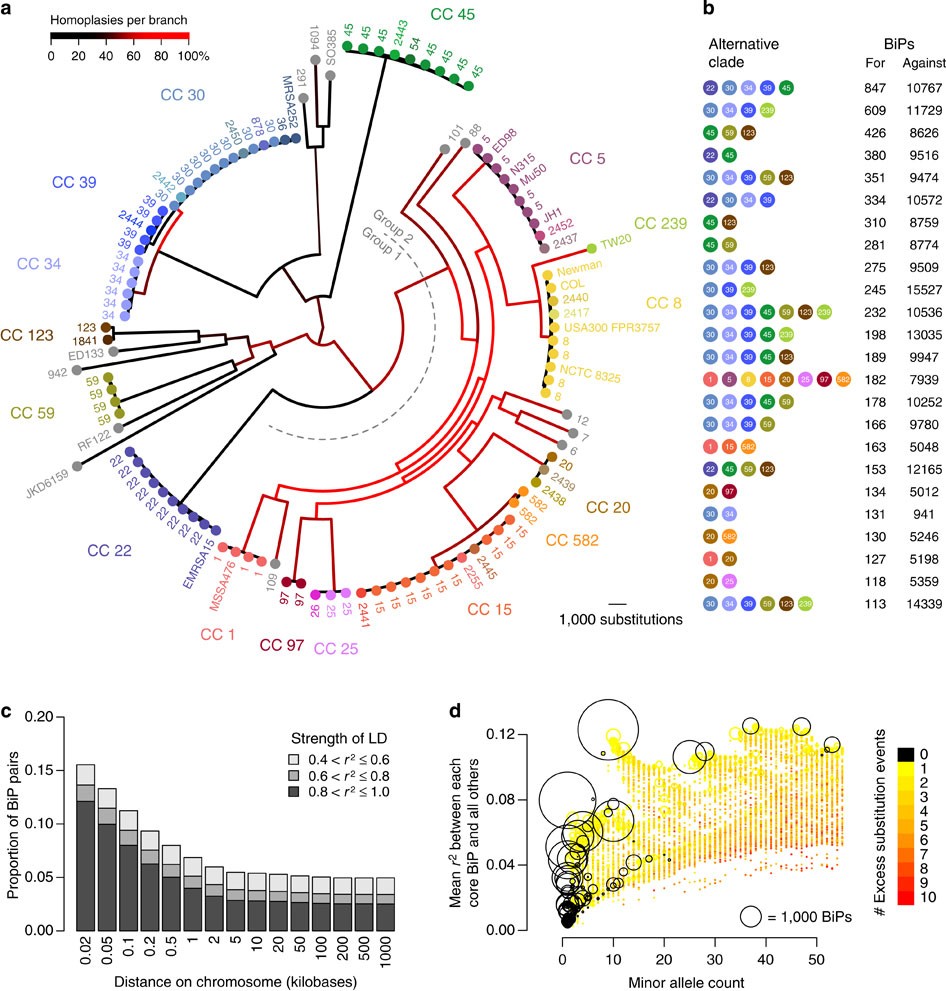
Evidence for a network-like organization across several conflicting phylogenetic signals was identified, and indeed several intragenic (within gene subdomains) recombination events were detected within and between CCs of MRSA strains. The alignment of the SpA sequences allowed the grouping of several isoforms as a result of non-randomly distributed amino acid variations, located in two groups of polymorphic sites in the D to B and Xr(a) domains. However, evidence of group-specific structural arrangements reflecting alterations in specific residues with a potential impact on the pathogenicity of S. aureus was detected.
Conclusions
The detection of positive selection operating in spA combined with frequent non-synonymous mutations, domain duplication, and frequent intragenic recombination events represent important mechanisms acting in the evolutionary adaptive mechanism that promotes the genetic plasticity of spA. These findings argue that crucial allelic forms correlated with pathogenicity can be identified by sequence analysis allowing the design of more robust schemes.
Electronic Supplementary Material
The online version of this article (doi:10.1186/s12866-016-0757-9) contains supplemental material, which is available to authorized users.
Keywords:
Staphylococcus aureus, staphylococcal protein A, recombination, molecular evolution, spA typing, virulence factor

